Rare Disease
Shedding Light on the Undiagnosed by GC Genome.
Enable a more complete story for rare disease patients with cutting edge genomic technologies.
What is rare disease testing?
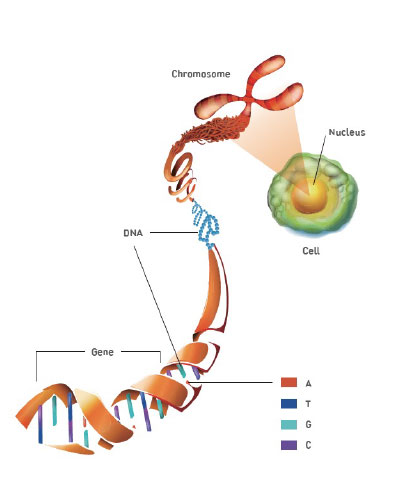
Most essential DNA coding for genetic information in humans is located at the nucleus of the cells. Genetic information is coded in a sequence of A, G, C, T nucleotides, and in human DNA there are approximately 3 billion sequences making up the genetic information.
Hereditary diseases are caused by a complete or partial change in these genetic sequences. The changes can be a mutation in one or multiple genes, mutation combined with environmental factors, or abnormalities in the number and structure of the chromosome.
There are approximately 7,000 types of rare diseases, with around 80% of them being hereditary and 50% affecting children. About 30% of patients with rare diseases may die before the age of 5. Accurate genetic diagnosis is essential for rare disease patients to receive appropriate treatment and management. Through GC Genome’s rare disease testing, we can find the possible cause (genetic variants).
Which Test Should I Select?
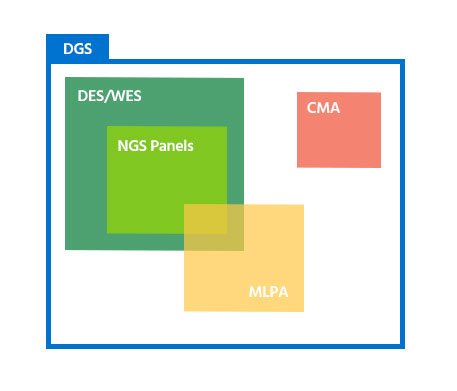
In order to successfully diagnose hereditary diseases, it is essential to select the most appropriate test based on types and sizes of the causal genetic variants.
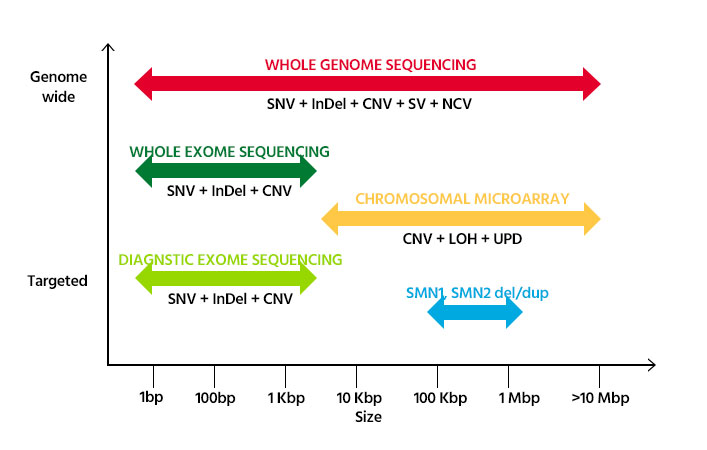
- SNV(Single Nucleotide Variant): A DNA sequence variation that occurs when a single nucleotide (adenine, thymine, cytosine, or guanine) in the genome
sequence is altered. - InDel (Insertion & Deletion): Additions or deletions of one or more nucleotides in a DNA sequence.
- CNV (Copy Number Variant): Sections of the genome are repeated and the number of repeats in the genome varies between individuals.
- SV (Structural Variant): Generally defined as a region of DNA approximately 1 kb and larger in size and can include inversions and balanced translocations.
- NCV (Non-Coding Variant): A variant in noncoding DNA can turn on a gene and cause a protein to be produced in the wrong place or at the wrong time.
- LOH (Loss of Heterozygosity): Genetic abnormality in diploid organisms in which one copy of an entire gene and its surrounding chromosomal region are lost.
- UPD (Uniparental disomy): Receiving two copies of a chromosome, or of part of a chromosome, from one parent and no copy from the other.
